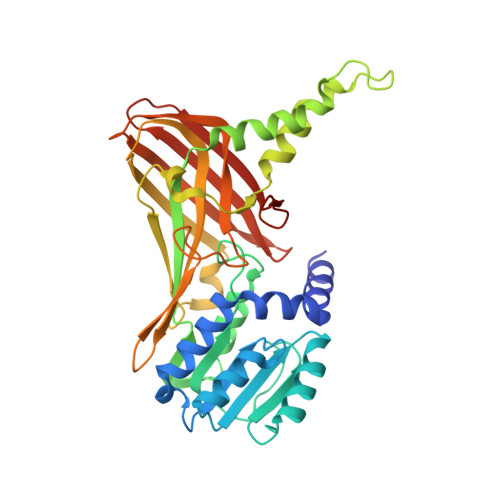
Functional insights from structures of coactivator-associated arginine methyltransferase 1 domains.
Troffer-Charlier, N., Cura, V., Hassenboehler, P., Moras, D., Cavarelli, J.(2007) EMBO J 26: 4391-4401
- PubMed: 17882262
- DOI: https://doi.org/10.1038/sj.emboj.7601855
- Primary Citation of Related Structures:
2OQB, 3B3F, 3B3G, 3B3J - PubMed Abstract:
Coactivator-associated arginine methyltransferase 1 (CARM1), a protein arginine methyltransferase recruited by several transcription factors, methylates a large variety of proteins and plays a critical role in gene expression. We report, in this paper, four crystal structures of isolated modules of CARM1. The 1.7 A crystal structure of the N-terminal domain of CARM1 reveals an unexpected PH domain, a scaffold frequently found to regulate protein-protein interactions in a large variety of biological processes. Three crystal structures of the CARM1 catalytic module, two free and one cofactor-bound forms (refined at 2.55 A, 2.4 A and 2.2 A, respectively) reveal large structural modifications including disorder to order transition, helix to strand transition and active site modifications. The N-terminal and the C-terminal end of CARM1 catalytic module contain molecular switches that may inspire how CARM1 regulates its biological activities by protein-protein interactions.
- Département de Biologie et Génomique Structurales, IGBMC (Institut de Génétique et de Biologie Moléculaire et Cellulaire), UMR 7104 CNRS, U596 INSERM, ULP, Illkirch, France.
Organizational Affiliation: