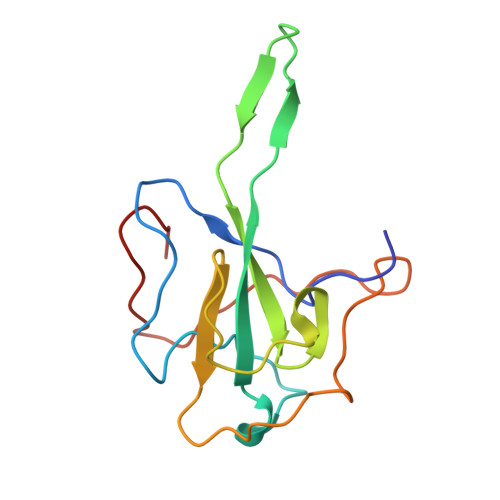
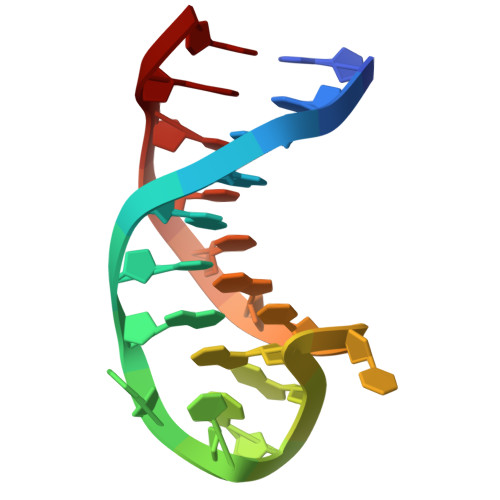
A compact stem-loop DNA aptamer targets a uracil-binding pocket in the SARS-CoV-2 nucleocapsid RNA-binding domain.
Esler, M.A., Belica, C.A., Rollie, J.A., Brown, W.L., Moghadasi, S.A., Shi, K., Harki, D.A., Harris, R.S., Aihara, H.(2024) Nucleic Acids Res 52: 13138-13151
- PubMed: 39380503
- DOI: https://doi.org/10.1093/nar/gkae874
- Primary Citation of Related Structures:
8TFD - PubMed Abstract:
SARS-CoV-2 nucleocapsid (N) protein is a structural component of the virus with essential roles in the replication and packaging of the viral RNA genome. The N protein is also an important target of COVID-19 antigen tests and a promising vaccine candidate along with the spike protein. Here, we report a compact stem-loop DNA aptamer that binds tightly to the N-terminal RNA-binding domain of SARS-CoV-2 N protein. Crystallographic analysis shows that a hexanucleotide DNA motif (5'-TCGGAT-3') of the aptamer fits into a positively charged concave surface of N-NTD and engages essential RNA-binding residues including Tyr109, which mediates a sequence-specific interaction in a uracil-binding pocket. Avid binding of the DNA aptamer allows isolation and sensitive detection of full-length N protein from crude cell lysates, demonstrating its selectivity and utility in biochemical applications. We further designed a chemically modified DNA aptamer and used it as a probe to examine the interaction of N-NTD with various RNA motifs, which revealed a strong preference for uridine-rich sequences. Our studies provide a high-affinity chemical probe for the SARS-CoV-2 N protein RNA-binding domain, which may be useful for diagnostic applications and investigating novel antiviral agents.
- Department of Biochemistry, Molecular Biology and Biophysics, University of Minnesota, Minneapolis, MN 55455, USA.
Organizational Affiliation: