Fe(II)-2-oxoglutarate-dependent pseudomonal iron uptake factor C in complex with malate
Alshref, F.M., Allen, M.D., Sanguankiattichai, N., Zaborskyte, G., Schofield, C.J.To be published.
Experimental Data Snapshot
Starting Model: in silico
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| PKHD-type hydroxylase PiuC | 225 | Pseudomonas aeruginosa | Mutation(s): 0 Gene Names: piuC, PA4515 EC: 1.14.11 | 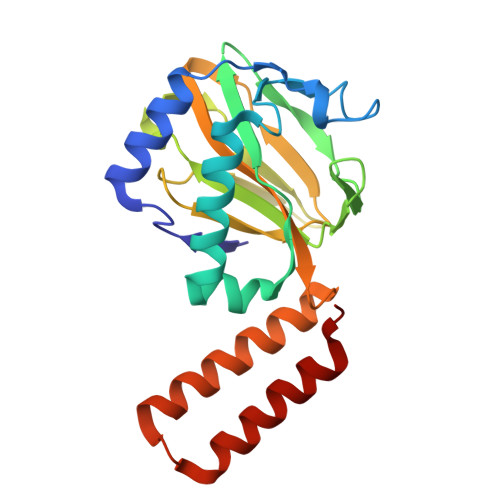 | |
UniProt | |||||
Find proteins for Q9HVQ7 (Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1)) Explore Q9HVQ7 Go to UniProtKB: Q9HVQ7 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9HVQ7 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| MLT (Subject of Investigation/LOI) Query on MLT | G [auth A] J [auth B] L [auth C] N [auth D] Q [auth E] | D-MALATE C4 H6 O5 BJEPYKJPYRNKOW-UWTATZPHSA-N |  | ||
| PEG Query on PEG | I [auth A] K [auth B] M [auth C] P [auth D] R [auth E] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| GOL Query on GOL | H [auth A], O [auth D], T [auth F] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 80.843 | α = 62.89 |
| b = 80.912 | β = 62.9 |
| c = 88.701 | γ = 60.03 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| DIALS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Saudi Ministry of Education | Saudi Arabia | DM8000S4747 |