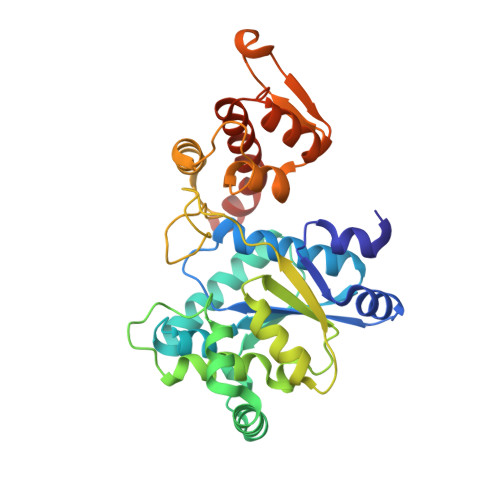
Structural plasticity of an aminoacyl-tRNA synthetase active site
Turner, J.M., Graziano, J., Spraggon, G., Schultz, P.G.(2006) Proc Natl Acad Sci U S A 103: 6483-6488
- PubMed: 16618920
- DOI: https://doi.org/10.1073/pnas.0601756103
- Primary Citation of Related Structures:
1ZH0, 2AG6 - PubMed Abstract:
Recently, tRNA aminoacyl-tRNA synthetase pairs have been evolved that allow one to genetically encode a large array of unnatural amino acids in both prokaryotic and eukaryotic organisms. We have determined the crystal structures of two substrate-bound Methanococcus jannaschii tyrosyl aminoacyl-tRNA synthetases that charge the unnatural amino acids p-bromophenylalanine and 3-(2-naphthyl)alanine (NpAla). A comparison of these structures with the substrate-bound WT synthetase, as well as a mutant synthetase that charges p-acetylphenylalanine, shows that altered specificity is due to both side-chain and backbone rearrangements within the active site that modify hydrogen bonds and packing interactions with substrate, as well as disrupt the alpha8-helix, which spans the WT active site. The high degree of structural plasticity that is observed in these aminoacyl-tRNA synthetases is rarely found in other mutant enzymes with altered specificities and provides an explanation for the surprising adaptability of the genetic code to novel amino acids.
- Department of Chemistry and The Skaggs Institute for Chemical Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: