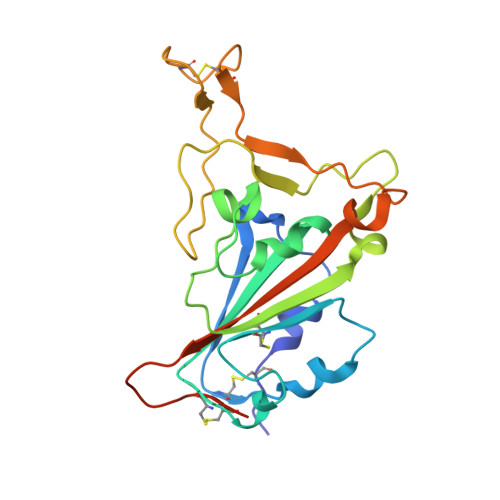
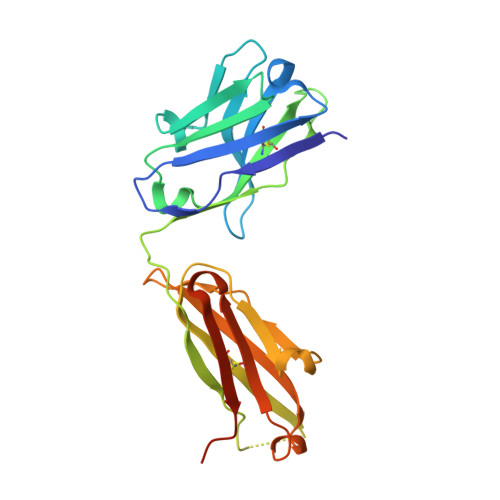
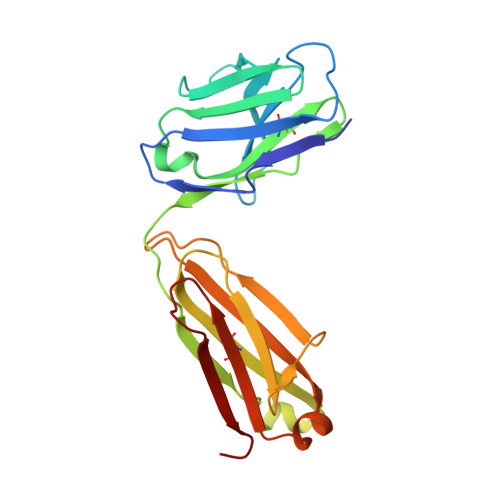
Broad sarbecovirus neutralization by combined memory B cell antibodies to ancestral SARS-CoV-2.
Planchais, C., Fernandez, I., Chalopin, B., Bruel, T., Rosenbaum, P., Beretta, M., Dimitrov, J.D., Conquet, L., Donati, F., Prot, M., Porrot, F., Planas, D., Staropoli, I., Guivel-Benhassine, F., Baquero, E., van der Werf, S., Haouz, A., Simon-Loriere, E., Montagutelli, X., Maillere, B., Rey, F.A., Guardado-Calvo, P., Nozach, H., Schwartz, O., Mouquet, H.(2024) iScience 27: 110354-110354
- PubMed: 39071888
- DOI: https://doi.org/10.1016/j.isci.2024.110354
- Primary Citation of Related Structures:
8QH0, 8QH1 - PubMed Abstract:
Antibodies play a pivotal role in protecting from SARS-CoV-2 infection, but their efficacy is challenged by the continuous emergence of viral variants. In this study, we describe two broadly neutralizing antibodies cloned from the memory B cells of a single convalescent individual after infection with ancestral SARS-CoV-2. Cv2.3194, a resilient class 1 anti-RBD antibody, remains active against Omicron sub-variants up to BA.2.86. Cv2.3132, a near pan-Sarbecovirus neutralizer, targets the heptad repeat 2 membrane proximal region. When combined, Cv2.3194 and Cv2.3132 form a complementary SARS-CoV-2 neutralizing antibody cocktail exhibiting a local dose-dependent synergy. Thus, remarkably robust neutralizing memory B cell antibodies elicited in response to ancestral SARS-CoV-2 infection can withstand viral evolution and immune escape. The cooperative effect of such antibody combination may confer a certain level of protection against the latest SARS-CoV-2 variants.
- Institut Pasteur, Université Paris Cité, INSERM U1222, Humoral Immunology Unit, 75015 Paris, France.
Organizational Affiliation: