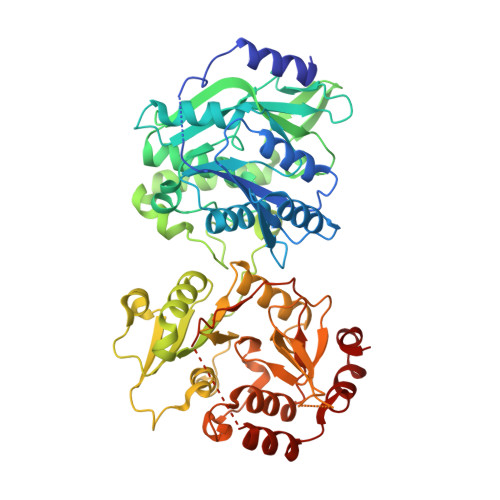
Structural and Functional Analysis of Heparosan Synthase 2 from Pasteurella multocida (PmHS2) to Improve the Synthesis of Heparin.
Stancanelli, E., Krahn, J.A., Viverette, E., Dutcher, R., Pagadala, V., Borgnia, M.J., Liu, J., Pedersen, L.C.(2024) ACS Catal 14: 6577-6588
- PubMed: 39990868
- DOI: https://doi.org/10.1021/acscatal.4c00677
- Primary Citation of Related Structures:
8VH7, 8VH8, 8VIW - PubMed Abstract:
Heparin is a widely used drug to treat thrombotic disorders in hospitals. Heparosan synthase 2 from Pasteurella multocida (PmHS2) is a key enzyme used for the chemoenzymatic synthesis of heparin oligosaccharides. It has both activities: glucosaminyl transferase activity and glucuronyl transferase activity; however, the mechanism to carry out the glyco-oligomerization is unknown. Here, we report crystal structures of PmHS2 constructs with bound uridine diphosphate (UDP) and a cryo-EM structure of PmHS2 in complex with UDP and a heptasaccharide (NS 7-mer) substrate. Using a LC-MC analytical method, we discovered the enzyme displays both a two-step concerted oligomerization mode and a distributive oligomerization mode depending on the non-reducing end of the starting oligosaccharide primer. Removal of 7 amino acid residues from the C-terminus results in an enzymatically active monomer instead of dimer and loses the concerted oligomerization mode of activity. In addition, the monomer construct can transfer N-acetyl glucosamine at a substrate concentration that is ∼7-fold higher than wildtype enzyme. It was also determined that an F529A mutant can transfer an N-sulfo glucosamine (GlcNS) saccharide from a previously inactive UDP-GlcNS donor. Performing the glyco-transfer reaction at a high substrate concentration and the capability of using unnatural donors are desirable to simplify the chemoenzymatic synthesis to prepare heparin-based therapeutics.
- Division of Chemical Biology and Medicinal Chemistry, Eshelman School of Pharmacy, University of North Carolina, Chapel Hill, North Carolina, USA.
Organizational Affiliation: