Integrating immune library probing with structure-based computational design to develop potent neutralizing nanobodies against emerging SARS-CoV-2 variants.
Cerdan, L., Silva, K., Rodriguez-Martin, D., Perez, P., Noriega, M.A., Esteban Martin, A., Gutierrez-Adan, A., Margolles, Y., Corbera, J.A., Martin-Acebes, M.A., Garcia-Arriaza, J., Fernandez-Recio, J., Fernandez, L.A., Casasnovas, J.M.(2025) MAbs 17: 2499595-2499595
- PubMed: 40329514
- DOI: https://doi.org/10.1080/19420862.2025.2499595
- Primary Citation of Related Structures:
9FC2 - PubMed Abstract:
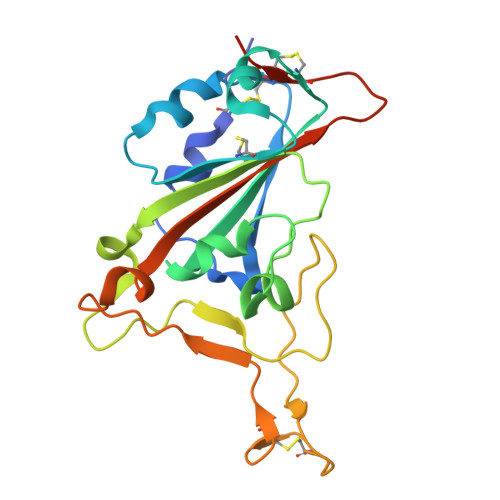
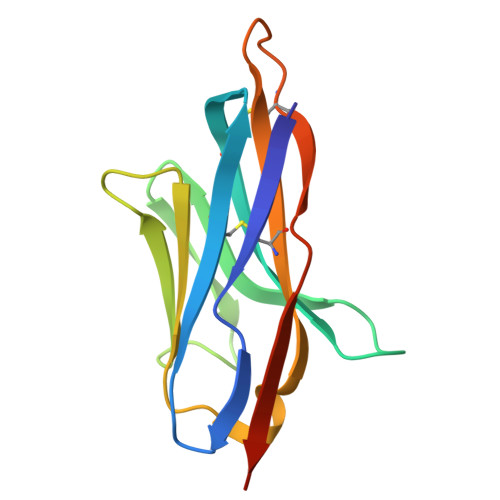
To generate antibodies (Abs) against SARS-CoV-2 emerging variants, we integrated multiple tools and engineered molecules with excellent neutralizing breadth and potency. Initially, the screening of an immune library identified a nanobody (Nb), termed Nb4, specific to the receptor-binding domain (RBD) of the Omicron BA.1 variant. A Nb4-derived heavy chain antibody (hcAb4) recognized the spike (S) of the Wuhan, Beta, Delta, Omicron BA.1, and BA.5 SARS-CoV-2 variants. A high-resolution crystal structure of the Nb4 variable (VHH) domain in complex with the SARS-CoV-2 RBD (Wuhan) defined the Nb4 binding mode and interface. The Nb4 VHH domain grasped the RBD and covered most of its outer face, including the core and the receptor-binding motif (RBM), which was consistent with hcAb4 blocking RBD binding to the SARS-CoV-2 receptor. In mouse models, a humanized hcAb4 showed therapeutic potential and prevented the replication of SARS-CoV-2 BA.1 virus in the lungs of the animals. In vitro , hcAb4 neutralized Wuhan, Beta, Delta, Omicron BA.1, and BA.5 viral variants, as well as the BQ.1.1 subvariant, but showed poor neutralization against the Omicron XBB.1.5. Structure-based computation of the RBD-Nb4 interface identified three Nb4 residues with a reduced contribution to the interaction with the XBB.1.5 RBD. Site-saturation mutagenesis of these residues resulted in two hcAb4 mutants with enhanced XBB.1.5 S binding and virus neutralization, further improved by mutant Nb4 trimers. This research highlights an approach that combines library screening, Nb engineering, and structure-based computational predictions for the generation of SARS-CoV-2 Omicron-specific Abs and their adaptation to emerging variants.
- Department of Microbial Biotechnology, Consejo Superior de Investigaciones Científicas (CNB-CSIC), Madrid, Spain.
Organizational Affiliation: