Structure of minimal CRISPR polymerase bound to ApNHpp
Doherty, E.E., Bolling, C.B., Wilcox, X.E., Doudna, J.A.To be published.
Experimental Data Snapshot
Starting Model: in silico
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| MCpol domain-containing protein | A [auth B], B [auth A] | 125 | Escherichia coli | Mutation(s): 0 Gene Names: DU321_04150 | 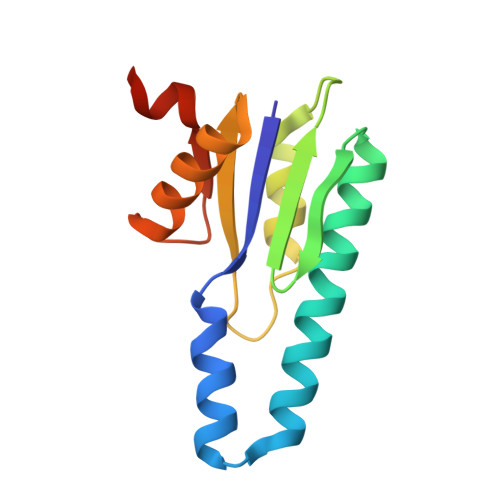 |
UniProt | |||||
Find proteins for A0A426EXS8 (Escherichia coli) Explore A0A426EXS8 Go to UniProtKB: A0A426EXS8 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A426EXS8 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ZAN (Subject of Investigation/LOI) Query on ZAN | C [auth B], E [auth A] | 5'-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]amino}phosphoryl]adenosine C10 H17 N6 O12 P3 WUPFAKUEHUVIDK-KQYNXXCUSA-N |  | ||
| MG (Subject of Investigation/LOI) Query on MG | D [auth B], F [auth A] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 49.135 | α = 90 |
| b = 56.769 | β = 90 |
| c = 93.705 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Howard Hughes Medical Institute (HHMI) | United States | -- |
| National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) | United States | F32GM153031 |